Table of contents:
Introduction.
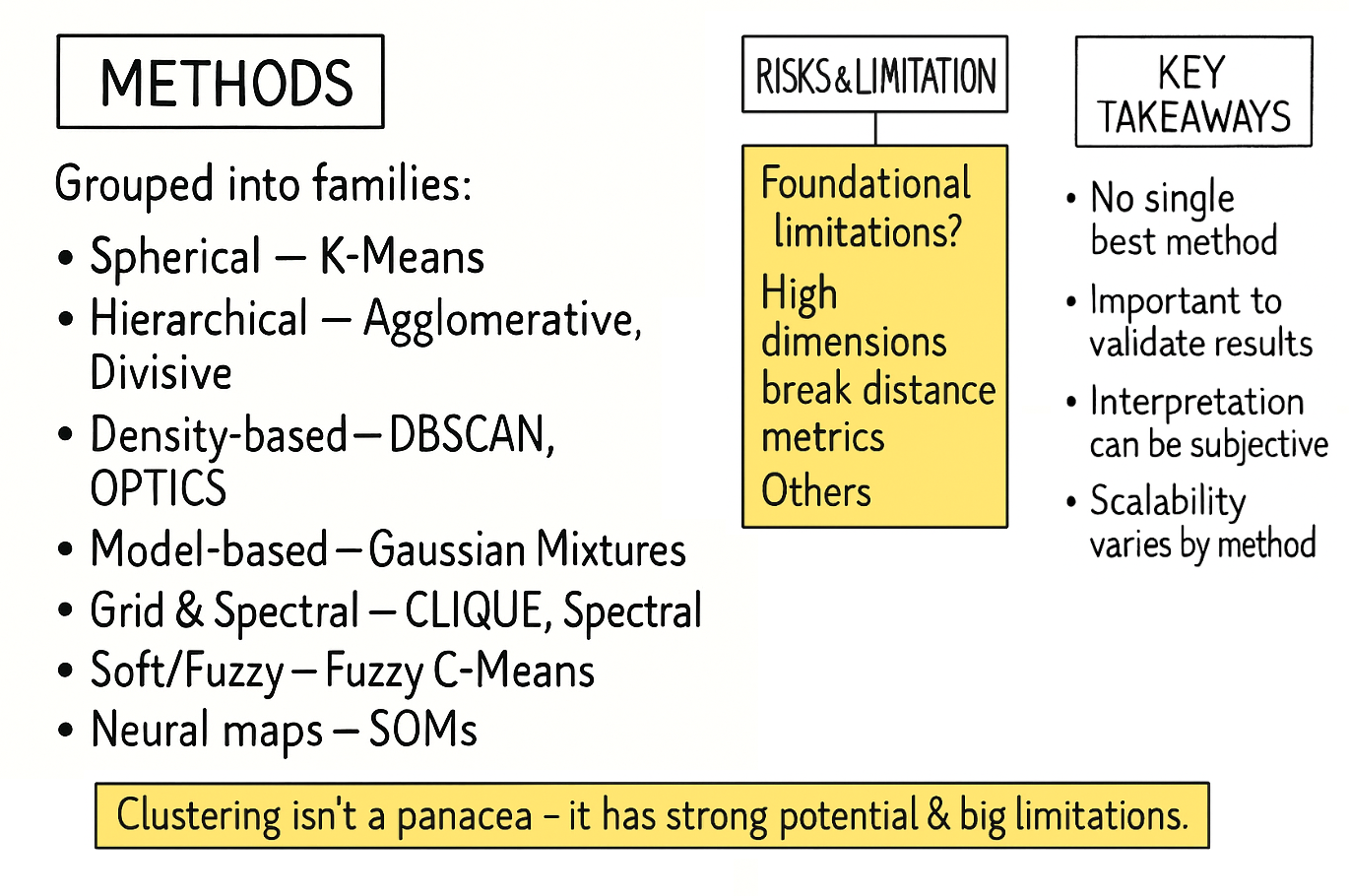
Risk and model limitations.
Spherical approaches.
K-Means.
K-Medoids.
Hierarchical approaches.
Density-based techniques.
DBSCAN.
OPTICS.
Model-based methods.
Grid-based & spectral methods.
CLIQUE.
Spectral clustering.
Fuzzy clustering.
Neural network-based methods.
Introduction
Alright, let’s establish first principles. Before deploying capital into algorithmic strategies, one must confront the paradigm shift that distinguishes durable firms from those erased by structural blind spots: financial markets are not monolithic stochastic processes but non-stationary systems governed by latent regime dynamics. This is not heuristic philosophy—it’s an empirical reality quantified through decades of regime-dependent risk premia and structural break analysis.
Every junior quant, armed with asymptotic theorems and clean data matrices, begins with the same intellectual shit: the assumption that markets exhibit global ergodicity. You construct elegant factor models—momentum, mean reversion, cointegration—optimized on historical time series under the illusion of stationarity. For a finite horizon, these models perform beautifully, their Sharpe ratios inflated by periods of regime persistence. Your backtests become exercises in overfitting, mistaking coincident stability for invariant truth.
This is the danger of deterministic thinking in a probabilistic domain. Your strategy thrives in a local equilibrium, calibrated to a specific regime’s statistical fingerprints—low realized volatility, persistent auto-correlation, stable cross-asset dependencies. But markets are adaptive systems. The moment regime boundaries shift—triggered by macroeconomic thresholds, liquidity phase transitions, or endogenous feedback loops—your model’s assumptions collapse. A momentum strategy tuned to trending markets withers under mean-reverting volatility spikes; a statistical arbitrage framework predicated on tight spreads disintegrates during liquidity droughts. The mathematics remain valid within the regime, but the system itself has mutated.
The epiphany arrives via PnL volatility: markets are better modeled as a mixture of distributions than a single stochastic process. Regime identification becomes the critical challenge. Formally, a regime is a metastable state characterized by distinct joint probability distributions over asset returns, volatility, correlation matrices, and microstructure signals. These states—think the "Complacent Bull" (low vol, positive drift), "Panicked Bear" (high vol, negative skew), and "Sideways Chop" (structural breakouts failing due to range-bound noise)—exhibit path dependency and memory effects.
Legacy models fail because they ignore regime segmentation, akin to applying linear regression to a piecewise function. The solution lies in unsupervised learning: algorithms that detect regime boundaries without labeled data. We are going to start by something simple like clustering techniques. Features might include realized volatility surfaces, cross-asset correlation eigenvalues, orderbook imbalance metrics, or entropy measures of liquidity concentration.
Clustering isn’t a panacea. It demands rigorous validation via silhouette scores, gap statistics, or out-of-sample stability tests. More critically, it requires domain-specific feature engineering to capture regime-defining characteristics. Seasoned practitioners know that regime detection is inherently recursive: today’s clusters inform tomorrow’s risk models, which refine the features used to identify the next regime shift.
If you are curious, check this:
There are no universal solutions, only frameworks for dynamic adaptation. The algorithms we’ll explore are not tools but lenses: each revealing partial truths about the market’s ever-shifting anatomy. Your survival hinges on switching these lenses faster than the system changes its shape. Let’s begin.
Risk and model limitations
Clustering is an essential technique for exploratory data analysis, allowing us to uncover latent structures in unlabeled data. However, it is crucial to recognize that a clustering algorithm is not an oracle that reveals pre-ordained truths. It is a mathematical procedure that optimizes a specific objective, and its results are highly sensitive to initial assumptions, algorithmic choices, and human interpretation. Understanding its limitations is the key to using it effectively.
The risks can be broadly categorized into three categories:
Foundational risks → Flaws in the premise:
These risks arise before any algorithm is run. They are fundamental to how the problem is framed.
The entire output of a clustering analysis hinges on the definition of "similarity" or "distance." This choice is made based on assumptions. An algorithm has no inherent understanding of context; it simply processes the mathematical representation of the features you provide. Choosing irrelevant or poorly scaled features will lead to mathematically valid but practically meaningless clusters.
As the number of features used to describe each data point increases, the concept of distance becomes less meaningful. In a high-dimensional space, the distance between any two points can appear to be almost equal. This makes it incredibly difficult for algorithms that rely on distance metrics—like K-Means or DBSCAN—to differentiate between a point's "close" and "distant" neighbors. The algorithm may find patterns, but these are often just artifacts of the vast, sparse space, not genuine structures in the data.
Price is not static, it evolves. A clustering model built on data from last year may be a poor representation of the current reality. This "concept drift" means that clusters are not permanent structures but transient states, and models must be regularly re-evaluated and retrained to remain relevant.
Algorithmic risks → Bias of the tools:
Every clustering algorithm has an intrinsic bias regarding the types of structures it is designed to find. Using the wrong tool for the data's underlying geometry is a common mistake.
Different algorithms are biased towards finding clusters of specific shapes. K-Means, for instance, will always try to find spherical, roughly equal-sized clusters because it works by minimizing variance around a central point. If the true clusters in your data are long and thin, or intertwined like crescent moons, K-Means will fail, incorrectly splitting them apart or merging them. DBSCAN can find arbitrary shapes but assumes all meaningful clusters have a similar density, which may not be the case. The algorithm will always return an answer, but that answer will be distorted if the data's true shape doesn't match the algorithm's bias.
Most clustering algorithms require the quant to set crucial parameters beforehand. The most famous is the number of clusters,
K, in K-Means. How do you know the correct number of clusters in advance? While methods exist to guide this choice—like the Elbow method—they are often ambiguous. Other algorithms are sensitive to different parameters, like the neighborhood radius—eps—in DBSCAN. A small change in these initial settings can lead to vastly different clustering outcomes, making the results seem arbitrary and unstable if the parameters cannot be robustly justified.Some algorithms are highly sensitive to outliers or noise. Since methods like K-Means attempt to assign every single point to a cluster, a few extreme outliers can significantly skew the position of a cluster's center, thereby corrupting the entire group. While some algorithms like DBSCAN have an explicit notion of "noise," those that don't can produce misleading results.
Interpretation risks → Fallacies in the conclusion:
Once the algorithm produces an output, the final and most critical set of risks emerges from human interpretation.
A clustering algorithm will always find clusters, even in perfectly random data. The human tendency is to seek patterns and assign narratives to these groupings. Just because you find a cluster and can describe its members' common characteristics does not mean the cluster represents a meaningful, underlying phenomenon. It might simply be a statistical coincidence. Attributing causality or deep significance to a mathematically derived group without external validation is a dangerous leap of faith.
It is easy to create a clustering model that perfectly partitions the data you used to build it. However, the true test of a model is how well it generalizes to new, unseen data. A model that is too complex or too finely tuned to the initial dataset has likely overfit the noise and random quirks present in that sample. When new data arrives, this overfit model will fail to assign it to the correct clusters, making it useless for any predictive or real-time application.
The boundaries between clusters are often fuzzy. A data point near a boundary can be re-assigned to a different cluster if the data is slightly perturbed or the model is re-run. This instability can make it difficult to rely on the classification for any critical downstream task. Furthermore, while we can mathematically describe a cluster by the average values of its features, this does not always translate to a simple, human-understandable concept.
The devil was in the execution. Clustering promised to be our compass, yet its limitations emerged like landmines in the path of progress. Further we will see how to avoid them.
Spherical approaches
My first instinct was to start simple. The most intuitive idea of clustering is simply to partition the data into a pre-specified number of groups, K. This leads us straight to the workhorse of clustering: K-Means.
K-Means
The concept is beautifully straightforward. Imagine you want to group all the trading days from the last five years into three regimes (K=3): "Bullish," "Bearish," and "Neutral." K-Means tries to find the best three central points—called centroids—to represent these regimes.
The algorithm is iterative:
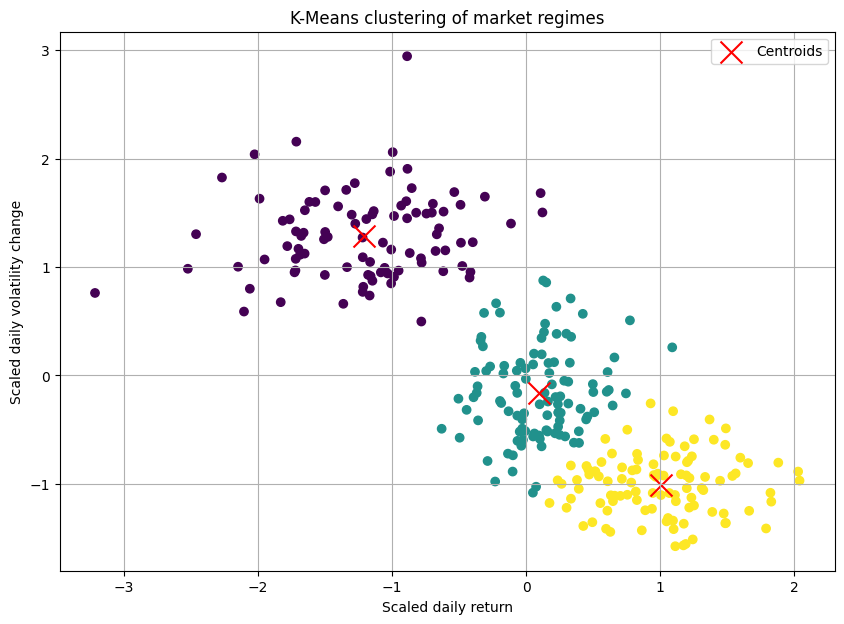
Initialization: First, it randomly places K centroids in your data space. In our case, imagine a 2D plot where the X-axis is the daily return of the S&P 500 and the Y-axis is the daily change in the VIX—volatility index. We'd just plop three points down randomly on this chart.
Assignment step: It goes through every single trading day and assigns it to the nearest centroid. The nearness is typically measured by standard Euclidean distance.
Update step: After assigning all the days, it recalculates the position of each centroid to be the mean—the average—of all the data points assigned to it.
Repeat: It repeats the assignment and update steps until the centroids stop moving, meaning a stable solution has been found.
The objective function that K-Means tries to minimize is the sum of squared distances from each data point xi to its assigned cluster's centroid μj. This is called the within-cluster sum of squares.
Where Cj is the set of all points in cluster j.
Let's simulate some market data. We'll create three distinct regimes:
Positive returns, low volatility change—bull quiet.
Negative returns, high volatility change—bear volatile.
Near-zero returns, moderate volatility change—sideways chop.
import numpy as np
from sklearn.cluster import KMeans
from sklearn.preprocessing import StandardScaler
# Generate synthetic market data
# We create data representing daily (Return, Volatility Change)
np.random.seed(42)
# Regime 1: Bull Quiet
bull_quiet = np.random.randn(100, 2) * [0.5, 0.5] + [1.0, -1.0]
# Regime 2: Bear Volatile
bear_volatile = np.random.randn(100, 2) * [0.6, 0.8] + [-1.2, 3.0]
# Regime 3: Sideways Chop
sideways_chop = np.random.randn(100, 2) * [0.3, 0.7] + [0.1, 0.5]
market_data = np.vstack([bull_quiet, bear_volatile, sideways_chop])
# Preprocessing
# It's crucial to scale data for distance-based algorithms like K-Means
scaler = StandardScaler()
scaled_data = scaler.fit_transform(market_data)
# Apply K-Means
kmeans = KMeans(n_clusters=3, random_state=42, n_init='auto')
clusters = kmeans.fit_predict(scaled_data)
centroids = kmeans.cluster_centers_The resulting plot would show our three blobs of data points, each colored differently, with a red 'X' marking the calculated center of each cluster. It looked beatufully artificial! Okay, we had successfully partitioned our market days.
Strengths:
K-Means is computationally efficient, making it suitable for large datasets.
Its logic is intuitive, making it a great starting point.
Pitfalls and our "oh, crap" moment:
The initial success quickly faded when we applied it to more complex data. The pitfalls became glaringly obvious.
We have to specify the number of clusters, K, beforehand. How do we know if there are 3, 4, or 10 market regimes? Choosing the wrong K leads to nonsensical groupings. We can use methods like the "Elbow Method," but often the "elbow" i ambiguous, leaving the choice subjective.
The initial random placement of centroids can lead to different final clusters. Running the algorithm multiple times with different random starts is a must.
K-Means implicitly assumes that clusters are spherical and evenly sized—this is a killer. Financial data is rarely so well-behaved. What if a "panic" regime is a long, thin, oddly shaped cluster? K-Means would fail spectacularly to capture it. It draws a circle where it should be drawing a snake.
A single extreme outlier event—like news—could drag a centroid way off course, corrupting an entire cluster.
To combat the outlier problem, we briefly experimented with K-Medoids—PAM - Partitioning Around Medoids. Instead of using the mean which is sensitive to outliers as the center, K-Medoids uses a medoid—an actual data point from the cluster—as the center.
K-Medoids
From a methodological standpoint, K-Medoids is a direct and robust alternative to K-Means. The primary limitation of K-Means stems from its use of the centroid—the mean of the points in a cluster—as the cluster prototype. The mean is a statistical measure that is notoriously sensitive to outliers. A single, aberrant data point can significantly skew the centroid, pulling it away from what we would intuitively consider the cluster's "center."
K-Medoids addresses this deficiency by a simple but key change: instead of a centroid, it uses a medoid. A medoid is constrained to be an actual data point from the dataset. It is defined as the point within a cluster that has the minimum average dissimilarity to all other points in that same cluster. By constraining the center to be an existing point, the algorithm becomes substantially more robust to the influence of extreme outliers and noise.
The most common algorithm for K-Medoids is PAM—Partitioning Around Medoids. Its objective is to minimize the sum of dissimilarities between the points and their respective cluster's medoid. However, it is much more computationally expensive and still suffered from the "K" and shape assumption problems. Our simple partitioning approach was too rigid for the fluid, strangely shaped reality of the market.
The algorithmic procedure PAM can be summarized as follows:
Initialization: Select K points from the dataset X to serve as the initial medoids.
Assignment: Assign each data point to the cluster corresponding to the nearest medoid. The total cost is the sum of the distances of all points to their medoid.
Update: For each cluster, consider swapping its medoid with a non-medoid point from the same cluster. Calculate the change in the total cost for each potential swap. If a swap would reduce the total cost, perform the swap that yields the greatest cost reduction.
Iteration: Repeat steps 2 and 3 until the medoids no longer change, indicating that the algorithm has converged to a local minimum.
The objective function of the K-Medoids algorithm is to find the set of medoids M and the corresponding partition C that minimizes the total within-cluster dissimilarity. This is expressed as:
Let's dissect this formula:
The inner summation calculates the sum of distances from every point xj within a single cluster Ci to that cluster's medoid, mi. This value represents the "cost" or total dissimilarity for that specific cluster.
The outer summation sums up the costs of all k individual clusters. The result is the total cost for the entire clustering configuration defined by the set of medoids M.
The minimization indicates that we are searching for the specific set of k medoids, chosen from the original dataset X, that results in the minimum possible value for this total cost function.
In essence, the formula seeks the most centrally located data points to act as exemplars for each cluster, where "centrality" is defined by the minimization of the sum of distances to all other points in that cluster.
Unlike K-Means, which minimizes the sum of squared Euclidean distances to an artificial centroid, the K-Medoids objective function typically uses the sum of absolute distances or other dissimilarity measures. This lack of squaring is a key reason for its increased robustness to outliers, as extreme distances are not amplified quadratically.
Let’s code it:
import numpy as np
from sklearn.cluster import KMeans
from sklearn.metrics import pairwise_distances
# Simplified K-Medoids (PAM) implementation
def kmedoids_simple(X, n_clusters, max_iter=100):
"""
A simplified implementation of the K-Medoids (PAM) algorithm.
Args:
X (np.ndarray): The input data, shape (n_samples, n_features).
n_clusters (int): The number of clusters (K).
max_iter (int): The maximum number of iterations.
Returns:
tuple: A tuple containing:
- labels (np.ndarray): Cluster labels for each point.
- medoid_centers (np.ndarray): The final medoid points.
"""
# 1. Initialize: Randomly select K data points as the initial medoids.
n_samples = X.shape[0]
initial_medoid_indices = np.random.choice(n_samples, n_clusters, replace=False)
medoids = X[initial_medoid_indices, :]
# Placeholder for the previous medoids to check for convergence
old_medoids = np.zeros_like(medoids)
for i in range(max_iter):
# 2. Assignment Step: Assign each point to the nearest medoid.
distances = pairwise_distances(X, medoids, metric='euclidean')
labels = np.argmin(distances, axis=1)
# 3. Update Step: For each cluster, find the point that minimizes the
# sum of distances within that cluster (the new medoid).
new_medoids = np.zeros_like(medoids)
for k in range(n_clusters):
# Get all points belonging to the current cluster
cluster_points = X[labels == k]
# If a cluster is empty, which can happen with bad initializations,
# re-assign a random point to it.
if len(cluster_points) == 0:
new_medoids[k, :] = X[np.random.choice(n_samples, 1), :]
continue
# Calculate the sum of distances from each point in the cluster
# to all other points in the same cluster.
intra_cluster_distances = pairwise_distances(cluster_points, metric='euclidean')
sum_of_distances = np.sum(intra_cluster_distances, axis=1)
# The point with the minimum sum of distances is the new medoid.
new_medoid_index_in_cluster = np.argmin(sum_of_distances)
new_medoids[k, :] = cluster_points[new_medoid_index_in_cluster, :]
# Check for convergence: If the medoids have not changed, stop.
if np.all(new_medoids == medoids):
break
medoids = new_medoids
return labels, medoids
# 1. Generate synthetic data
# This section is identical to your original code.
np.random.seed(42)
# Create three distinct clusters
X1 = np.random.randn(50, 2) + np.array([0, 0])
X2 = np.random.randn(50, 2) + np.array([8, 8])
X3 = np.random.randn(50, 2) + np.array([0, 8])
# Add two significant outliers
outliers = np.array([
[15, 15], # Outlier affecting cluster X2
[-5, 15] # Outlier affecting cluster X3
])
# Combine into a single dataset
X = np.vstack([X1, X2, X3, outliers])
# 2. Apply clustering algorithms
# K-Means Algorithm (unchanged)
kmeans = KMeans(n_clusters=3, random_state=42, n_init=10)
kmeans_labels = kmeans.fit_predict(X)
kmeans_centers = kmeans.cluster_centers_
# K-Medoids algorithm (using our self-contained function)
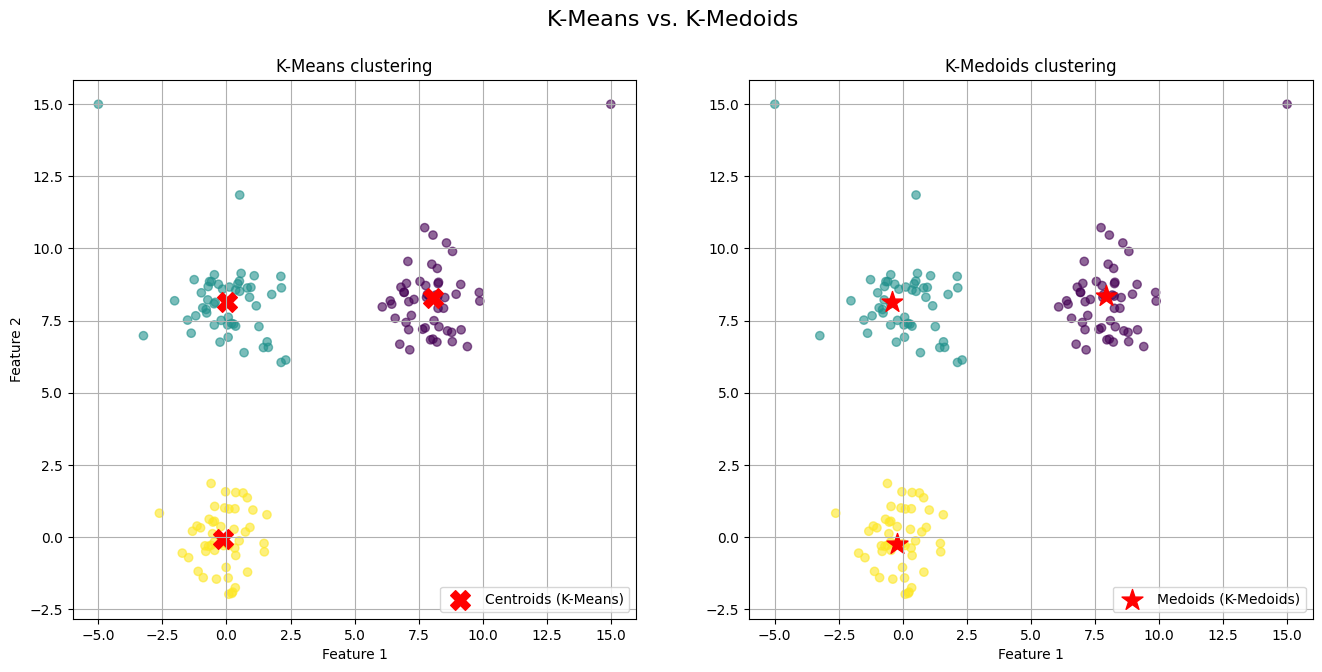
kmedoids_labels, kmedoids_centers = kmedoids_simple(X, n_clusters=3)The generated visualization provides a clear and direct comparison of the two algorithms' behaviors.
On the left plot (K-Means) we observe the red 'X' markers representing the centroids. Notice the centroids for the top-right and top-left clusters. They have been noticeably "pulled" away from the dense core of their respective clusters and shifted towards the outliers we introduced. The centroid is no longer a faithful representative of its cluster's central tendency; it is a mathematical average biased by extreme values.
On the right plot (K-Medoids) we observe the red star markers representing the medoids. The medoids for all three clusters are positioned squarely within the dense, central region of their assigned points. Because a medoid must be one of the actual data points, it cannot be an artificial point in space like a centroid. The algorithm selects the most "central" existing point, effectively ignoring the pull of distant outliers.
Now that we know the main strength, but pitfalls continue. It still suffers from the "K" and shape assumption problems.
Hierarchical approaches
Our failure with K-Means leads to a shift in perspective. Maybe market regimes aren't like separate, disconnected islands. Maybe they're related, nested within each other, like a family tree. For instance, a broad high volatility environment could contain sub-regimes like high-vol trend and high-vol chop. This line of thinking leads us to hierarchical clustering.
This method doesn't produce a single flat partition of the data but rather a hierarchy of clusters. There are two main approaches:
Agglomerative (bottom-up): This is the more common approach. It starts with every single data point in its own cluster. Then, in each step, it merges the two closest clusters until only one giant cluster—containing all the data—remains.
Divisive (top-down): It starts with all data points in a single cluster and recursively splits it into smaller clusters.
The real magic of hierarchical clustering is the dendrogram. It's a tree diagram that visualizes the entire hierarchy of merges—or splits. You can then "cut" the dendrogram at a certain height to get a specific number of clusters, but its real power is in showing the relationships.
The key component is the linkage criterion, which defines how to measure the distance between two clusters (not just two points). Common linkage criteria include:
Single linkage: Distance is the minimum distance between any two points in the two clusters. It can find long, thin clusters but is sensitive to noise.
\(d_{\text{single}}(C_A, C_B) = \min_{a \in C_A, b \in C_B} d(a, b)\)Complete linkage: Distance is the maximum distance between any two points. It produces more compact clusters.
\(d_{\text{complete}}(C_A, C_B) = \max_{a \in C_A, b \in C_B} d(a, b)\)Average linkage: Distance is the average distance between all pairs of points. It's a balance between the two.
\(d_{\text{average}}(C_A, C_B) = \frac{1}{|C_A| \cdot |C_B|} \sum_{a \in C_A} \sum_{b \in C_B} d(a, b)\)Ward's linkage: This is often the default. It merges the two clusters that result in the minimum increase in the total within-cluster variance.
\(\text{WCSS}(C_k) = \sum_{\boldsymbol{x}_i \in C_k} \|\boldsymbol{x}_i - \boldsymbol{\mu}_k\|^2\)
Besides the choice of the distance function d(⋅,⋅) is a critical modeling decision:
Let's apply this to the same market data:
import scipy.cluster.hierarchy as sch
from sklearn.cluster import AgglomerativeClustering
# Using Ward's linkage method
dendrogram = sch.dendrogram(sch.linkage(scaled_data, method='ward'))
# Perform agglomerative clustering
# Based on the dendrogram, we choose 3 clusters
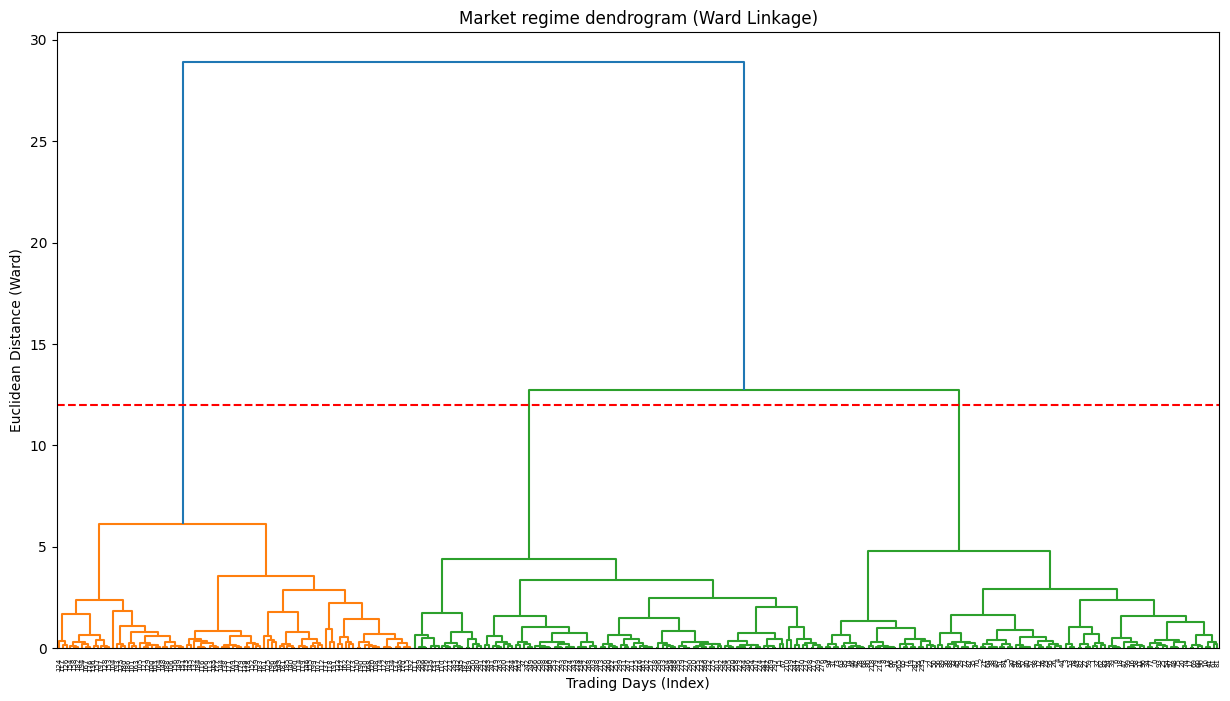
agg_cluster = AgglomerativeClustering(n_clusters=3, metric='euclidean', linkage='ward')
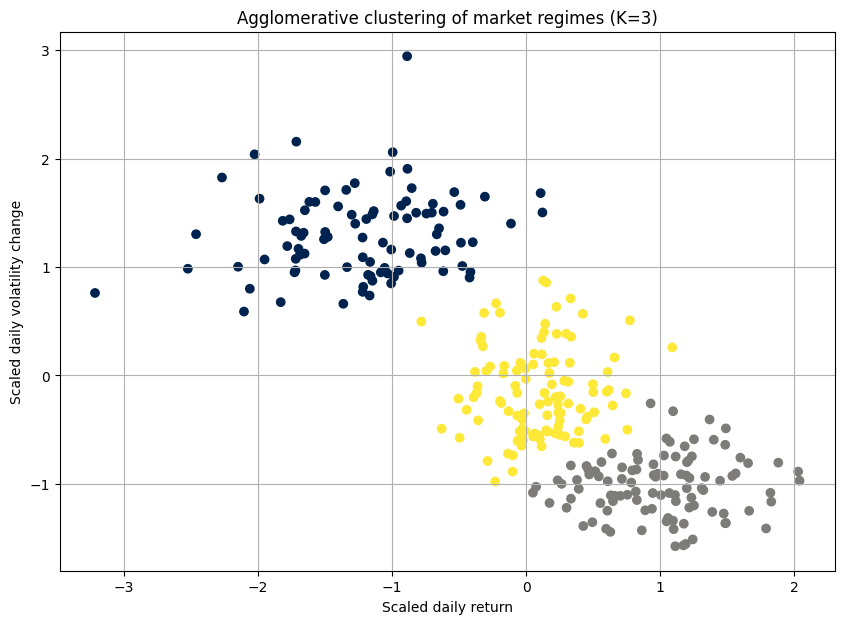
clusters = agg_cluster.fit_predict(scaled_data)The dendrogram is the key output here. It's a beautiful, complex tree. The y-axis represents the distance at which clusters were merged. By drawing a horizontal line, we can select a number of clusters. For example, where our red dashed line cuts the tree, it crosses three vertical lines, suggesting that three is a reasonable number of clusters.
Strengths:
The number of clusters is not required beforehand. The dendrogram provides a rich visualization of possibilities.
It doesn't just partition; it shows how clusters relate to one another, which can be very insightful for understanding market dynamics.
It can work with any distance metric, not just Euclidean.
Pitfalls and a new frustrations:
Hierarchical clustering felt like a more honest approach. But it wasn't a silver bullet.
The basic agglomerative algorithm has a time complexity of at least O(N2logN), where N is the number of data points. For our dataset of thousands of trading days, it was slow.
While we don't have to specify K upfront, we still have to decide where to "cut" the dendrogram. This decision remains subjective and can dramatically change the interpretation.
Once a merge—in agglomerative—or a split—in divisive—is made, it's final. An early mistake can't be corrected later in the process.
For a high-dimensional dataset, the 2D representation of the dendrogram can be misleading. The distances it portrays might not capture the true, complex relationships in the data.
I have found a way to map the market's family tree, but the map was hard to read and computationally expensive to draw.